Notebooks
Premium
Trends
BioTuring
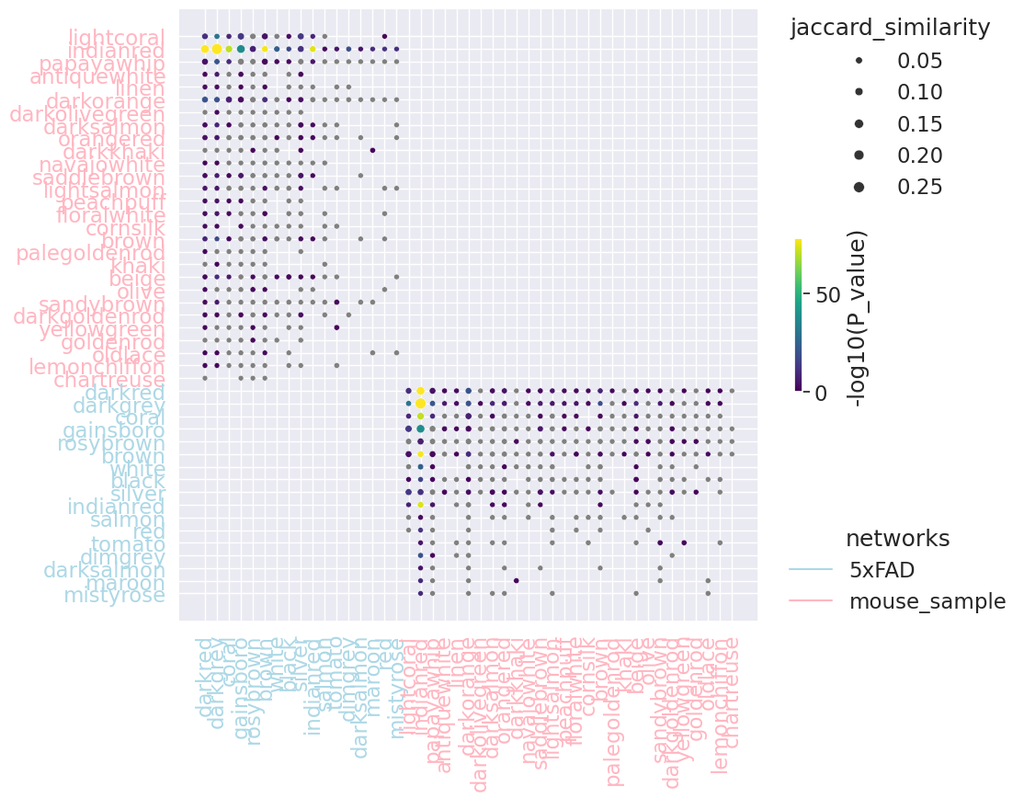
PyWGCNA is a Python library designed to do weighted correlation network analysis (WGCNA). It can be used for:
- Finding clusters (modules) of highly correlated genes
- For summarizing such clusters using the module eigengene
- For relating modules to one another and to external sample traits (using eigengene network methodology)
- For calculating module membership measures.
Users can also compare WGCNA networks from different datasets, or to external gene lists, to assess the conservation or functional enrichment of each module.