Notebooks
Premium
Trends
BioTuring
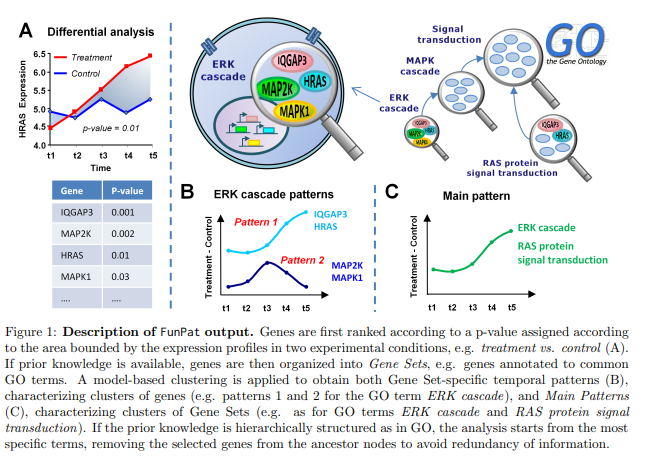
Dynamic expression data, nowadays obtained using high-throughput RNA sequencing (RNA-seq), are essential to monitor transient gene expression changes and to study the dynamics of their transcriptional activity in the cell or response to stimuli. FunPat is an R package designed to provide:
- a useful tool to analyze time series genomic data;
- a computational pipeline which integrates gene selection, clustering and functional annotations into a single framework to identify the main temporal patterns associated to functional groups of differentially expressed (DE) genes;
- an easy way to exploit different types of annotations from currently available databases (e.g. Gene Ontology) to extract the most meaningful information characterizing the main expression dynamics;
- a user-friendly organization and visualization of the outcome, automatically linking the DE genes and their temporal patterns to the functional information for an easy biological interpretation of the results.